Guest posting: Assoc. Prof. Josh Davis, Principal Research Fellow/NHMRC Career Development Fellow, Menzies School of Health Research, Senior Staff Specialist Infectious Diseases Physician, John Hunter Hospital, Conjoint Professor School of Medicine and Public Health, University of Newcastle
The diverse bacterial communities which live in our gastrointestinal tract (primary in the colon), are collectively known as the “gut microbiota” and their collective genes are termed the “microbiome“. A majority of the bacterial species cannot cultivated and require direct molecular techniques to ennumerate them. Consideration of the microbiome is extremely topical in many fields of health-related research. At the recent major annual meeting of the American Society for Microbiology (ASM Microbe), the gut microbiome had its own stream (no pun intended – and the colour coding wasn’t brown) within the conference program, containing scores of research presentations in this field. In fact, one conference delegate, science writer and microbiologist Ed Yong, insisted on introducing himself as “me and my multitudes” (referring to his microbiome) rather than just as a person (see his YouTube channel for some edifying entertainment).
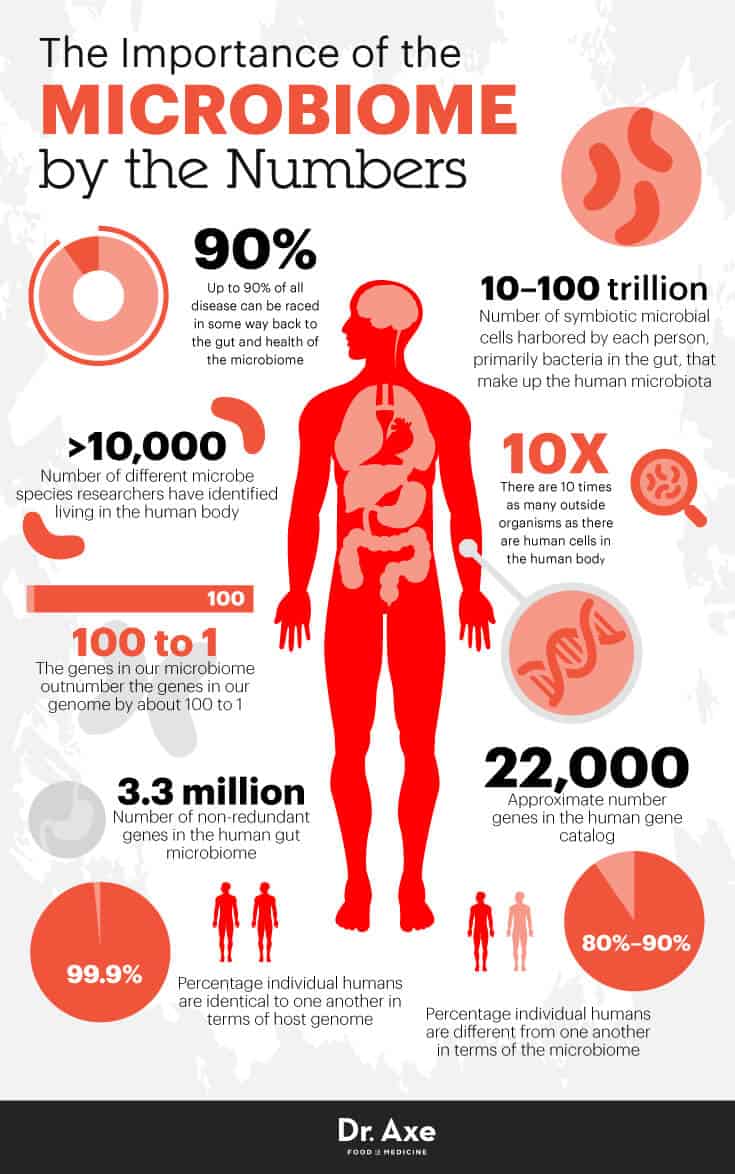
There are an estimated 3 million different genes in the average human’s microbiome, as compared with 20,000 genes in the human genome. This means that the metabolic activity of all of these microbial genes is vastly complex and could potentially influence processes ranging from digestion of food through oncogenesis to inflammation. Some examples from ASM Microbe 2017 include:
- A study presented by Dr Heidi Nelson, a surgeon from the Mayo Clinic in USA. They analysed the gut microbiome from 500 patients with colonic adenomatous polyps (the precursors of bowel cancer) and compared this with matched controls without colonic polyps. Those with the colonic polyps had more hydrogen sulphide-producing and pro-inflammatory microbes in their colons than the controls. Is this chicken or egg?
- The gut microbiome is associated with obesity. If you transplant the stool from an obese rat into a normal weight one, the recipient becomes obese. Dr Jun Chen, also from the Mayo clinic analysed the microbiome of a large number of healthy Americans, and reported that those with a BMI>30 have lower bacterial species diversity in their guts than those of healthy weight.
- Dr D Zeevi and team from the Weizman institute in Israel even proposed that the gut microbiome can be used to help estimate an individual’s response to a glycaemic food challenge. They have developed an algorithm incorporating gut microbiome data as well as demographics and dietary habits to create a personalised nutrition plan to avoid hyperglycaemia and/or diabetes. (ncbi.nlm.nih.gov/pubmed/26590418).
So, in the coming era of personalised medicine, we are going to need to think about not only our own personal genome, but that of our microbiome. Think about this next time you are planning to prescribe or take a course of antibiotics: antibiotics have a profound effect on our gut microbiome, which in turn relates to many of our physiological and pathological processes. . . .if we were considering using a drug that causes human genetic mutations, we would hesitate. Perhaps we should think in the same way about antibiotics.
This is such an interesting field! We will hear more about the effects of the gut microbiota on drug pharmacokinetics (and corresponding bioavailability) and thus efficacy/adverse effects, as well as the effects of drugs on the gut microbiome itself (e.g., PPIs and NSAIDs, statins etc).
LikeLike
Reblogged this on Microbiology and Infectious Diseases postgraduate teaching.
LikeLike